pgv-pmauve
pgv-pmauve is one of the CLI workflows in pyGenomeViz for
visualization of genome alignment results using progressiveMauve.
It can be used to visualize commonly conserved regions between all genomes.

Installation
Conda
conda install -c conda-forge -c bioconda pygenomeviz progressivemauve
Pip
pip install pygenomeviz
Additional installation of progressiveMauve is required. On Ubuntu, progressiveMauve can be installed with apt command.
sudo apt install progressivemauve
Docker
docker run -it --rm -p 8501:8501 ghcr.io/moshi4/pygenomeviz:latest pgv-pmauve -h
Usage
$ pgv-pmauve --help
usage: pgv-pmauve [options] seq1.gbk seq2.gbk seq3.gbk -o outdir
pyGenomeViz CLI workflow using progressiveMauve
positional arguments:
seqs Input genbank or fasta files
General Options:
-o , --outdir Output directory
--formats Output image format ('png'[*],'jpg','svg','pdf',`html`[*])
--reuse Reuse previous alignment result if available
-q, --quiet No print log on screen (default: OFF)
-v, --version Print version information
-h, --help Show this help message and exit
Figure Appearence Options:
--fig_width Figure width (Default: 15)
--fig_track_height Figure track height (Default: 1.0)
--track_align_type Figure tracks align type ('left'|'center'[*]|'right')
--feature_track_ratio Feature track ratio (Default: 0.25)
--show_scale_bar Show scale bar (Default: OFF)
--show_scale_xticks Show scale xticks (Default: OFF)
--curve Plot curved style link (Default: OFF)
--dpi Figure DPI (Default: 300)
--track_labelsize Track label size (Default: 20)
--scale_labelsize Scale label size (Default: 15)
--normal_link_color Normal link color (Default: 'grey')
--inverted_link_color Inverted link color (Default: 'red')
--refid Reference genome index (Default: 0)
--block_plotstyle Synteny block plot style ('box'[*]|'bigbox')
--block_cmap Synteny block colormap (Default: 'gist_rainbow')
[*] marker means the default value.
Multi Contigs Input Case
progressiveMauve cannot align for multiple contigs. Therefore, when multiple contigs are included in the input, a sequence that concatenates the multiple contigs into one is used as the input.
Colormap(--block_cmap) Option
User can arbitrarily choose the visualization colormap from
the various colormap types in matplotlib.
See Choosing Colormaps in Matplotlib for details.
Examples
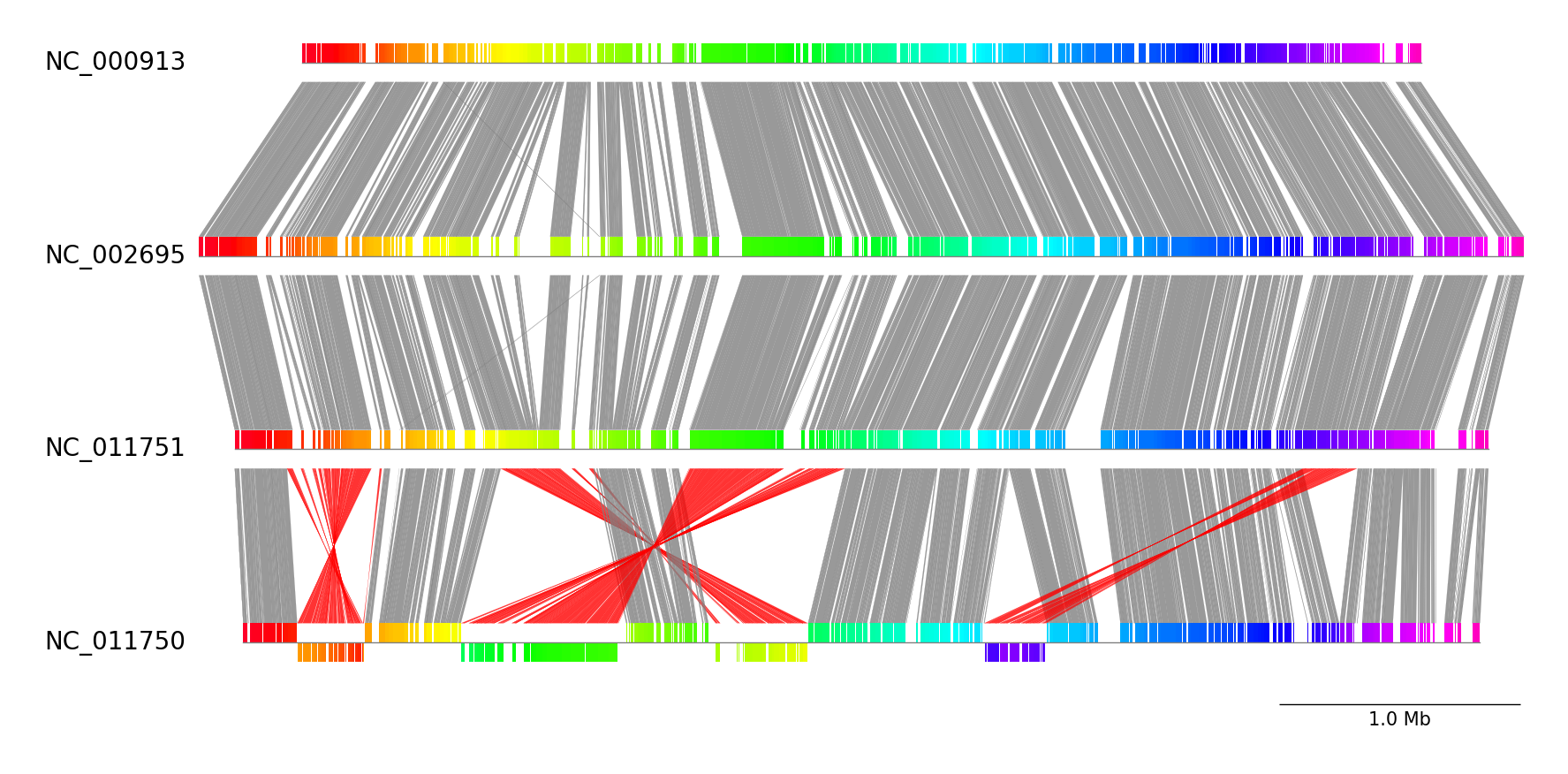
Example 1
Download example dataset:
pgv-download escherichia_coli
Run CLI workflow:
pgv-pmauve NC_000913.gbk.gz NC_002695.gbk.gz NC_011751.gbk.gz NC_011750.gbk.gz \
-o pgv-pmauve_example1 --show_scale_bar

Example 2
Download example dataset:
pgv-download mycoplasma_mycoides
Run CLI workflow:
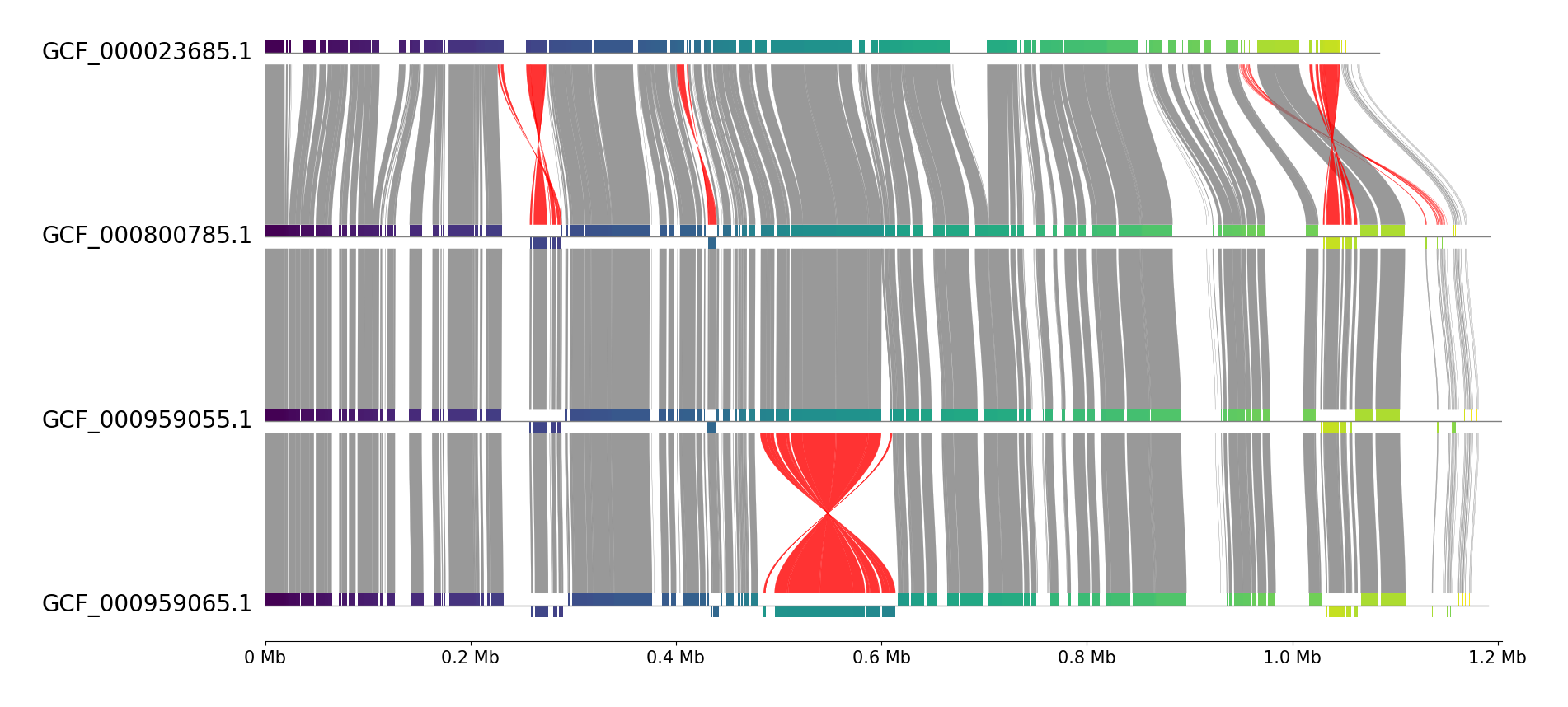
pgv-pmauve GCF_000023685.1.gbff GCF_000800785.1.gbff GCF_000959055.1.gbff GCF_000959065.1.gbff \
-o pgv-pmauve_example2 --feature_track_ratio 0.15 --block_cmap viridis --track_align_type left \
--show_scale_xticks --curve