pgv-mummer
pgv-mummer is one of the CLI workflows in pyGenomeViz for
visualization of genome alignment using MUMmer(nucmer, promer).

Installation
Conda
conda install -c conda-forge -c bioconda pygenomeviz mummer
Pip
pip install pygenomeviz
Additional installation of MUMmer is required. On Ubuntu, MUMmer can be installed with apt command.
sudo apt install mummer
Docker
docker run -it --rm -p 8501:8501 ghcr.io/moshi4/pygenomeviz:latest pgv-mummer -h
Usage
$ pgv-mummer --help
usage: pgv-mummer [options] seq1.gbk seq2.gbk seq3.gbk -o outdir
pyGenomeViz CLI workflow using MUMmer (nucmer, promer)
positional arguments:
seqs Input genbank files
General Options:
-o , --outdir Output directory
--formats Output image format ('png'[*],'jpg','svg','pdf',`html`[*])
--reuse Reuse previous alignment result if available
-q, --quiet No print log on screen (default: OFF)
-v, --version Print version information
-h, --help Show this help message and exit
MUMmer Alignment Options:
--seqtype Alignment sequence type ('nucleotide'[*]|'protein')
--threads Threads number (Default: MaxThread - 1)
--length_thr Length threshold to be plotted (Default: 0)
--identity_thr Identity threshold to be plotted (Default: 0)
Figure Appearence Options:
--fig_width Figure width (Default: 15)
--fig_track_height Figure track height (Default: 1.0)
--track_align_type Figure tracks align type ('left'|'center'[*]|'right')
--feature_track_ratio Feature track ratio (Default: 0.25)
--show_scale_bar Show scale bar (Default: OFF)
--show_scale_xticks Show scale xticks (Default: OFF)
--curve Plot curved style link (Default: OFF)
--dpi Figure DPI (Default: 300)
--track_labelsize Track label size (Default: 20)
--scale_labelsize Scale label size (Default: 15)
--normal_link_color Normal link color (Default: 'grey')
--inverted_link_color Inverted link color (Default: 'red')
--segment_space Track segment space ratio (Default: 0.02)
--feature_type2color Feature plot type & color (Default: ['CDS:orange'])
--pseudo_color Pseudo feature plot color (Default: 'lightgrey')
--feature_plotstyle Feature plot style ('[big]arrow'[*]|'[big]box'|'[big]rbox')
--feature_linewidth Feature line width (Default: 0.0)
--feature_labeltrack Feature label target track ('top'[*]|'all')
--feature_labeltype Feature label type ('product'|'gene'|'protein_id'|'None'[*])
--feature_labelsize Feature label size (Default: 8)
--cbar_width Colorbar width (Default: 0.01)
--cbar_height Colorbar height (Default: 0.2)
[*] marker means the default value.
--seqtype Option
--seqtype nucleotide: nucmer, --seqtype protein: promer
Examples
Example 1
Download example dataset:
pgv-download yersinia_phage
Run CLI workflow:
pgv-mummer NC_070914.gbk NC_070915.gbk NC_070916.gbk NC_070918.gbk \
-o pgv-mummer_example1 --show_scale_bar
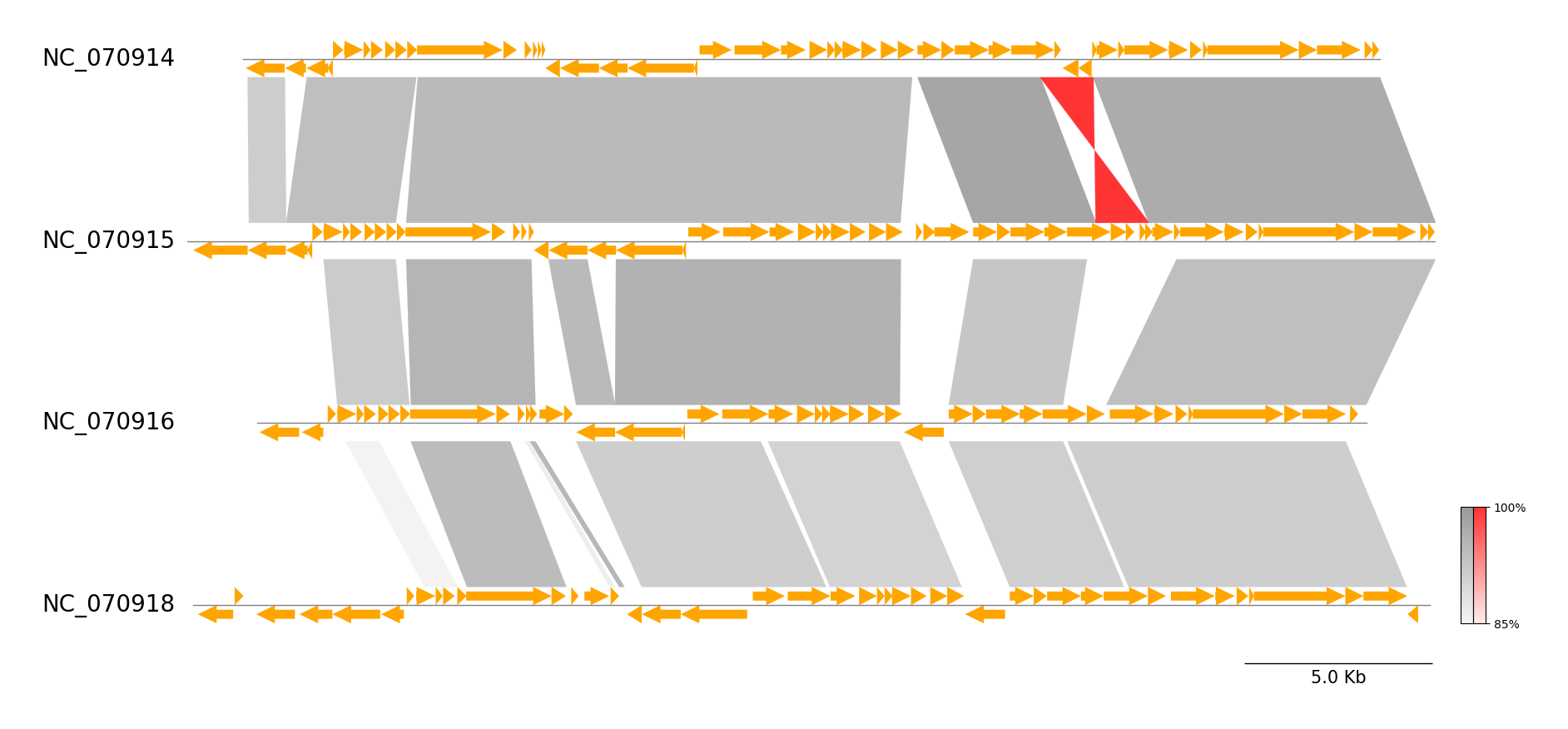
Example 2
Download example dataset:
pgv-download enterobacteria_phage
Run CLI workflow:
pgv-mummer NC_013600.gbk NC_016566.gbk NC_019724.gbk NC_024783.gbk NC_028901.gbk NC_031081.gbk \
-o pgv-mummer_example2 --seqtype protein --show_scale_bar --curve \
--feature_track_ratio 0.15 --fig_track_height 0.7 --feature_linewidth 0.5 --feature_plotstyle bigarrow \
--feature_type2color CDS:skyblue --normal_link_color chocolate --inverted_link_color limegreen
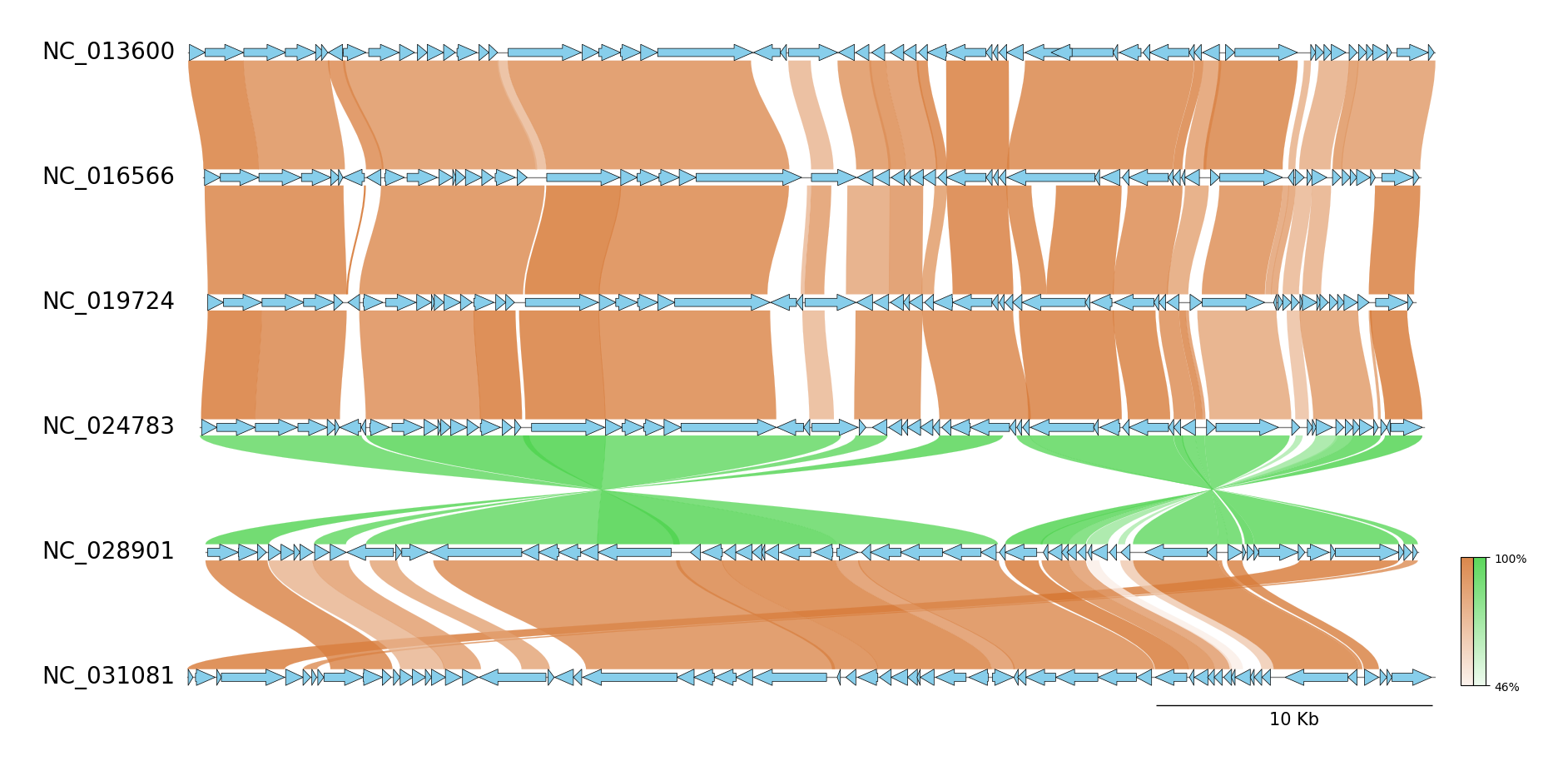
Example 3
Download example dataset:
pgv-download mycoplasma_mycoides
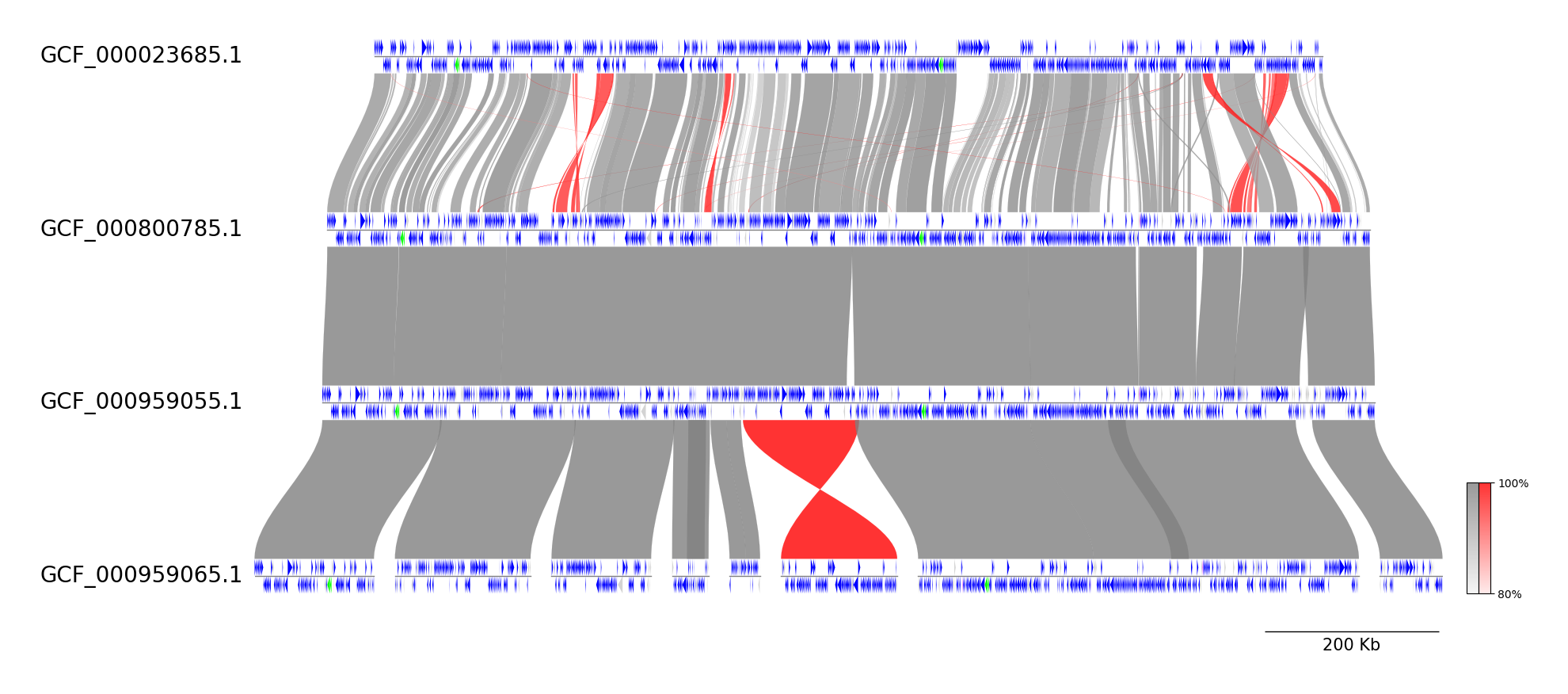
Run CLI workflow:
pgv-mummer GCF_000023685.1.gbff GCF_000800785.1.gbff GCF_000959055.1.gbff GCF_000959065.1.gbff \
-o pgv-mummer_example3 --show_scale_bar --curve \
--feature_type2color CDS:blue rRNA:lime tRNA:magenta
